Ancova
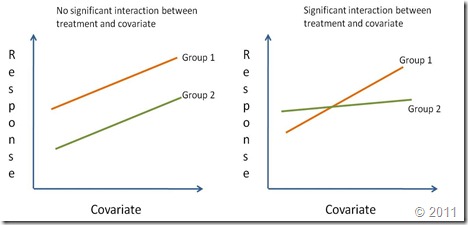
The analysis of covariance (ANCOVA) is used to compare two or more regression lines by testing the effect of a categorical factor on a dependent variable (y-var) while controlling for the effect of a continuous co-variable (x-var). When we want to compare two or more regression lines, the categorical factor splits the relationship between x-var and y-var into several linear equations, one for each level of the categorical factor.Regression lines are compared by studying the interaction of the categorical variable (i.e. treatment effect) with the continuous independent variable (x-var). If the interaction is significantly different from zero it means that the effect of the continuous covariate on the response depends on the level of the categorical factor. In other words, the regression lines have different slopes (Right graph on the figure below). A significant treatment effect with no significant interaction shows that the covariate has the same effect for all levels of the categorical factor. However, since the treatment effect is important, the regression lines although parallel have different intercepts. Finally, if the treatment effect is not significant nor its interaction with the covariate (but the coariate is significant), this means there is a single regression line. A reaction norm is used to graphically represent the possible outcomes of an ANCOVA.
For this example we want to determine how body size (snout-vent length) relates to pelvic canal width in both male and female alligators (data from Handbook of Biological Statistics). In this specific case, sex is a categorical factor with two levels (i.e. male and female) while snout-vent length is the regressor (x-var) and pelvic canal width is the response variable (y-var). The ANCOVA will be used to assess if the regression between body size and pelvic width are the comparable between the sexes.
Interpretation
An ANCOVA is able to test for differences in slopes and intercepts among regression lines. Both concepts have different biological interpretations. Differences in intercepts are interpreted as differences in magnitude but not in the rate of change. If we are measuring sizes and regression lines have the same slope but cross the y-axis at different values, lines should be parallel. This means that growth is similar for both lines but one group is simply larger than the other. A difference in slopes is interpreted as differences in the rate of change. In allometric studies, this means that there is a significant change in growth rates among groups.Slopes should be tested first, by testing for the interaction between the covariate and the factor. If slopes are significantly different between groups, then testing for different intercepts is somewhat inconsequential since it is very likely that the intercepts differ too (unless they both go through zero). Additionally, if the interaction is significant testing for natural effects is meaningless (see The Infamous Type III SS). If the interaction between the covariate and the factor is not significantly different from zero, then we can assume the slopes are similar between equations. In this case, we may proceed to test for differences in intercept values among regression lines.
Performing an ANCOVA
For an ANCOVA our data should have a format very similar to that needed for an Analysis of Variance. We need a categorical factor with two or more levels (i.e. sex factor has two levels: male and female) and at least one independent variable and one dependent or response variable (y-var).The preceding code shows the first six lines of the gator object which includes three variables: sex, snout and pelvic, which hold the sex, snout-vent size and the pelvic canal width of alligators, respectively. The sex variable is a factor with two levels, while the other two variables are numeric in their type.> head(gator) sex snout pelvic 1 male 1.10 7.62 2 male 1.19 8.20 3 male 1.13 8.00 4 male 1.15 9.60 5 male 0.96 6.50 6 male 1.19 8.17
We can do an ANCOVA both with the lm() and aov() commands. For this tutorial, we will use the aov() command due to its simplicity.
The previous code shows the ANCOVA model, pelvic is modeled as the dependent variable with sex as the factor and snout as the covariate. The summary of the results show a significant effect of snout and sex, but no significant interaction. These results suggest that the slope of the regression between snout-vent length and pelvic width is similar for both males and females.> mod1 <- aov(pelvic~snout*sex, data=gator) > summary(mod1) Df Sum Sq Mean Sq F value Pr(>F) snout 1 51.871 51.871 134.5392 8.278e-13 *** sex 1 2.016 2.016 5.2284 0.02921 * snout:sex 1 0.005 0.005 0.0129 0.91013 Residuals 31 11.952 0.386 --- Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
A second more parsimonious model should be fit without the interaction to test for a significant differences in the slope.
> mod2 <- aov(pelvic~snout+sex, data=gator)
> summary(mod2)
Df Sum Sq Mean Sq F value Pr(>F)
snout 1 51.871 51.871 138.8212 3.547e-13 ***
sex 1 2.016 2.016 5.3948 0.02671 *
Residuals 32 11.957 0.374
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
The second model shows that sex has a significant effect on the dependent variable which in this case can be interpreted as a significant difference in ‘intercepts’ between the regression lines of males and females. We can compare mod1 and mod2 with the anova() command to assess if removing the interaction significantly affects the fit of the model:The anova() command clearly shows that removing the interaction does not significantly affect the fit of the model (F=0.0129, p=0.91). Therefore, we may conclude that the most parsimonious model is mod2. Biologically we observe that for alligators, body size has a significant and positive effect on pelvic width and the effect is similar for males and females. However, we still don’t know how the slopes change.> anova(mod1,mod2) Analysis of Variance Table Model 1: pelvic ~ snout * sex Model 2: pelvic ~ snout + sex Res.Df RSS Df Sum of Sq F Pr(>F) 1 31 11.952 2 32 11.957 -1 -0.0049928 0.0129 0.9101
At this point we are going to fit linear regressions separately for males and females. In most cases, this should have been performed before the ANCOVA. However, in this example we first tested for differences in the regression lines and once we were certain of the significant effects we proceeded to fit regression lines.
To accomplish this, we are now going to sub-set the data matrix into two sets, one for males and another for females. We can do this with the subset() command or using the extract functions []. We will use both in the following code for didactic purposes:
Separate regression lines can also be fitted using the subset option within the lm() command, however we will use separate data frames to simplify the creation of graphs:> machos <- subset(gator, sex=="male") > hembras <- gator[gator$sex=='female',]
> reg1 <- lm(pelvic~snout, data=machos); summary(reg1)
Call:
lm(formula = pelvic ~ snout, data = machos)
Residuals:
Min 1Q Median 3Q Max
-0.85665 -0.40653 -0.08933 0.04518 1.57408
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) 0.4527 0.9697 0.467 0.647
snout 6.5854 0.8625 7.636 6.85e-07 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.7085 on 17 degrees of freedom
Multiple R-squared: 0.7742, Adjusted R-squared: 0.761
F-statistic: 58.3 on 1 and 17 DF, p-value: 6.846e-07
> reg2 <- lm(pelvic~snout, data=hembras); summary(reg2)
Call:
lm(formula = pelvic ~ snout, data = hembras)
Residuals:
Min 1Q Median 3Q Max
-0.69961 -0.19364 -0.07634 0.04907 1.15098
Coefficients:
Estimate Std. Error t value Pr(>|t|)
(Intercept) -0.2199 0.9689 -0.227 0.824
snout 6.7471 0.9574 7.047 5.8e-06 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.4941 on 14 degrees of freedom
Multiple R-squared: 0.7801, Adjusted R-squared: 0.7644
F-statistic: 49.67 on 1 and 14 DF, p-value: 5.797e-06
The regression lines indicate that males have a higher intercept (a=0.45) than females (a=-0.2199), which means that males are larger. We can now plot both regression lines as follows:
> plot(pelvic~snout, data=gator, type='n')
> points(machos$snout,machos$pelvic, pch=20)
> points(hembras$snout,hembras$pelvic, pch=1)
> abline(reg1, lty=1)
> abline(reg2, lty=2)
> legend("bottomright", c("Male","Female"), lty=c(1,2), pch=c(20,1) )
The resulting plot shows the regression lines for males and females on the same plot.Advanced
We can fit both regression models with a single call to the lm() command using the nested structure of snout nested within sex (i.e. sex/snout) and removing the single intercept for the model so that separate intercepts are fit for each equation.
> reg.todo <- lm(pelvic~sex/snout - 1, data=gator)
> summary(reg.todo)
Call:
lm(formula = pelvic ~ sex/snout - 1, data = gator)
Residuals:
Min 1Q Median 3Q Max
-0.85665 -0.33099 -0.08933 0.05774 1.57408
Coefficients:
Estimate Std. Error t value Pr(>|t|)
sexfemale -0.2199 1.2175 -0.181 0.858
sexmale 0.4527 0.8498 0.533 0.598
sexfemale:snout 6.7471 1.2031 5.608 3.76e-06 ***
sexmale:snout 6.5854 0.7558 8.713 7.73e-10 ***
---
Signif. codes: 0 ‘***’ 0.001 ‘**’ 0.01 ‘*’ 0.05 ‘.’ 0.1 ‘ ’ 1
Residual standard error: 0.6209 on 31 degrees of freedom
Multiple R-squared: 0.9936, Adjusted R-squared: 0.9928
F-statistic: 1213 on 4 and 31 DF, p-value: < 2.2e-16


in your example, you showed an ancova where the interaction term is not significant (ie best model has different intercepts but same slope)...yet you have plotted it with separate slopes. How would you plot it with different intercepts but the same slope? thanks.
ReplyDeleteTom,
ReplyDeleteIf you have estimated the common slope, then you can plot a differnt line for each group with different intercepts and the same slope using the abline() command
abline(a,b)
where a is the intercept for males or females and b is the common slope.
An interesting alternative to:
ReplyDeletereg.todo <- lm(pelvic~sex/snout - 1, data=gator)
is this formula:
reg.todo <- lm(pelvic~snout*sex, data=gator)
All of the coefficients will be the same for the two models (i.e. both fit a separate slope and intercept for M and F gators) but there are two differences, one superficial, one really interesting. The superficial one is the way the coefficients are presented. In your example, you get the true coefficients, in the second example (snout*sex) you'll get the coefficients for the first factor (females) and then the differences in the coefficient for the male factor. You can just add the male coefficients to the female ones so they're on the same scale.
The interesting thing in the summary of the second model is that the t score and p-value test different hypotheses than they do in the first model. In the first one they test whether or not the regression has a nonzero slope, in the second one, they test whether or not the slopes and intercepts from different factors are statistically significant! Nifty, right?
This comment has been removed by the author.
ReplyDeleteThanks James W. for your comment. I knew about lm(y~x*f) but I didn't know the differences regarding hypotheses testing. Thanks for the insight.
ReplyDeleteVery useful Campitor!!
ReplyDeleteJames W I did not know that neither! so interesting. I did it as individual regresions to get de R^2 and P of the individuals valors for each level of the factor.
Thank you- this was very helpful. If you have 3 factors (Treatments A,B,C) and you find that the slopes are not different among the treatments (no significant interaction- as in model 1), but that the intercepts are different (as in model 2). How do you test for which treatments are significantly different from each other? Thank you!
ReplyDeleteAR
This comment has been removed by the author.
DeleteHope this URL would be helpful. ####http://rcompanion.org/rcompanion/e_04.html####
DeleteAR - I've seen people use the Tukey HSD test to do this, and this should be okay because theoretically the different treatments/factors are independent, but I'm not sure. You could also bypass the formal test and simply present the intercepts and their standard error to demonstrate how they relate to each other.
ReplyDeleteThank you.
ReplyDeleteAR
Hi,
ReplyDeleteThank you for this guide. I am a bit confused that the slopes on the plot for male and female regression lines look the same. However, you say that the slops are totally different (one positive and one negative), but that does not seem to be the case. Am I misunderstanding something, or is this a typo?
Aras,
ReplyDeleteYou are right, that was a typo. the difference is in "intercepts" rather than slopes. Slopes are very similar between males and females indeed (6.5 and 6.7, respectively). the slopes are different and they indicate a change in the mean size of each group. In this case, males are significantly larger than females, as was shown by the ANOVA.
Thanks for clarifying that. So when you say "A second more parsimonious model should be fit without the interaction to test for a significant differences in the slope." should that also be intercept and not "slope"? I am not trying to be picky, its just that in my data I think there is a significant difference in intercept and not slope, and I am following your tutorial to prove it. Just want to cover all the bases.
ReplyDeleteCheers!
Aras,
ReplyDeleteNo, the interaction term test for differences in slopes. The difference in intercepts (or means) is tested by the natural factor (i.e. sex). If both regression lines have the same intercept, but dramatically different slopes (imagine two lines diverging from the same point on the y-axis), the interaction would be significant. You may use James W procedure (in the comments) to test for differences in intercepts. Hope this helps.
Thank you for your reply, its much more clear now.
ReplyDeleteHello Campitor,
ReplyDeletethank you very much for your easy to follow and excellently structured walk-through ANCOVA.
Just for curiosity. You determine the regression coefficients with subsets for "hembras" and "machos".
Is there no function in R which can calculate these parameters based on the model Y ~ X + A for all levels of A without the need of subsets?
Thanks a lot for your nice blog.
By the way... I could not find an "impressum". Is that by intention?
Sincerely
Karl-Heinz
Hi Karl,
ReplyDeletethanks for your comments. Regarding your first question, you can get parameters for all levels in two ways, the first one is described in the Advanced section. The other way is following James W instructions in the comments.
I'm not sure what you mean by "Impressum", if you mean a printer ready version of this topic, the reason it's not there is because I don't know how to do it. :).
Hi Campitor,
ReplyDeleteGreat tutorial! I have a question about adding a second covariate. Say that you want to look at the effect of an intervention. You have a pre-test score and a post-test score, two groups (treatment, controls). Normally, you would model it as:
post_test~pre_test + group
vs.
post_test~pre_test*group
But what if you wanted to add a second covariate. How do you factor out the influence of gender(M,F)? Does it look like this:
post_score~pre_score+group+gender
vs.
post_score~pre_score*group*gender
Hi,
ReplyDeletethanks a lot for your very helpful tutorial.
I wonder, how would you interpret a result that indicates that the slopes of males and females ARE different and, for example, cross over in the middle? What if the slopes are different BUT they will actually cross over at a point far beyond the range of your data?
Thanks
A really helpful tutorial, many thanks!
ReplyDeleteI may have missed something but I was wondering where the '(b ≈ 7.07; weighted average)' came from you speak of in the conclusion. I don't think this value is found in any of the outputs.
Cheers
Hello, this is a great tutorial! I used it a month ago for my own research. Now, I am wondering if I could use the figures and logic for a course that I am teaching. I'm wondering too if you made all those figures yourself for this post, so that I could attribute things correctly.
ReplyDeleteThanks, hopefully. Great job!
@Jeff Cardille
ReplyDeleteHi. I'm glad the tutorial was useful. You may definitely use the logic and images for your class. I did make all of them.
I'm going to post a new tutorial soon, but I'm still deciding on what specific topic. Be sure to keep posted.
Ok, thanks much! Your other postings are excellent as well by the way...
ReplyDeleteThank you so much. I search for R help a lot and never find such concise, well-written answers. I got everything on the first try and it actually looks the way you said it would....that never happens.
ReplyDeleteThank you so much. I search for R help a lot and never find such concise, well-written answers. I got everything on the first try and it actually looks the way you said it would....that never happens.
ReplyDeleteHi, this is a nice post.
ReplyDeleteI am wondering whether the non-significant intercepts (in reg1 and reg2) are indeed different. According to the average value and the sd of the value its seems that probably the "true" value include zero for both cases, so I do not get how it is possible to say that are different. Perhaps I am missing something.
Regards
Angel
Greetings, I am curious how you would modify this code to include an interaction model that is adjusted for other covariates?
ReplyDeleteGreetings, I am curious how you would modify this code to include an interaction model that is adjusted for other covariates?
ReplyDeleteIs there any way you can share the data? (gator) I would be very interesting in being able to replicate the analysis.
ReplyDeleteMany thanks,
Bernhard
This was incredibly useful, thank you!!! First time I've come across help with R that i actually understand. If you ever felt like writing something about how to look for statistical differences between nonlinear regressions that would be amazing :)
ReplyDeleteHi Rachel, thanks for your comments. I have not had much time to write in this blog, but I should probably get back to it this summer.
ReplyDeleteBy non-linear do you mean glm or really non-linear regression (like exponential or Gompertz?)
Camp
Hello,
ReplyDeleteI have a similar case, but my categorical factor has 4 treatments. The interaction is non significant, but the treatment effect and coavariate are significant. Does that means that the 4 treatments have different intercepts? Or it could be that just some of them have differences between each other? In that case, is it possible to do a contrast within intercepts?
Thanks a lot, your blog is very helpful!
Monica
Thank you for the great guide!
ReplyDeleteI have a simple question that might be a bit silly. Regarding scaling relationships, the common function used to describe how one trait scales with body size is: y = bx^a. Only when you log it, it becomes a linear relationship: logy=alogx+logb
My question is, when we are analyzing data for allometry using linear regression, do we have to log it? It seams so taking the equations into account.
And then when you used the log data for the analysis, the given intercept is a real b or a log(b)?
Thank you!
Thank you so much for the much needed tutorial.
ReplyDeleteI failed to transfer the points into the plot window with the given function. I got the following error message.
"Error in points(FIN$scale(CP), FIN$log(Cstock + 1), pch = 20) :
attempt to apply non-function"
Would you help me, please?
Thanks,
John
Thank you for this very helpful guide.
ReplyDeleteI have a similar situation where the interaction term is significant, but I have many treatments. Do you know of any way to test between the individual slopes applied to each treatment? I expect that there will be a few treatments which respond much more strongly to the covariate and would like to identify them statistically rather than visually.
HI, first of all, thank you very much for this tutorial, it is by far the most useful I've ever read. I have one doubt: in my study, I found no significant interaction effect, but the regression lines overlap.
ReplyDeleteFor instance in mod1 the effect of sex is significant, but the interaction effect is not
In mod2, the sex is still significant.
Then when I run anova(mod1,mod2), the F is non-significant, like in your example.
So, why my regression lines overlap?
Where did I go wrong? Is this normal?
How do you compare regression lines with nonparametric data? Can you still use ANCOVA?
ReplyDelete
ReplyDeleteParabéns pela receita, testei e aprovei. Tenho certeza que se você vendesse e usasse máquina de cartão para autônomo provavelmente aumentaria sua conversão.
Well Said, you have furnished the right information that will be useful to anyone at all time. Thanks for sharing your Ideas.
ReplyDeleteR Programming Online Training
Thank you. This is probably the clearest example i could find and made me understood the workings of ANCOVA in minutes. Thank you for the enlightenment!
ReplyDeleteI am extremely overwhelmed by the clarity and the conciseness of your presentation. It is super understandable. Thank you.
ReplyDeleteThank you, thank you for this post!!! :) Very well explained.
ReplyDeleteThank you for the information.
ReplyDeleteBest Training and Real Time Support
Oracle Osb Training
Power Bi Training
Amazing article. Your blog helped me to improve myself in many ways thanks for sharing this kind of wonderful informative blogs in live. I have bookmarked more article from this website. Such a nice blog you are providing ! Kindly Visit Us
ReplyDeleteRead more : R Programming institutes in Chennai | R Programming Training in Chennai
ReplyDeleteI think things like this are really interesting. I absolutely love to find unique places like this. It really looks super creepy though!! big data training in Velachery | Hadoop Training in Chennai | big data Hadoop training and certification in Chennai | Big data course fees
Paylaştığınız bilgiler, çok iyi ve ilginç. Bu makaleyi okuduğum için şanslıyım
ReplyDeletecửa lưới chống muỗi
lưới chống chuột
cửa lưới dạng xếp
cửa lưới tự cuốn
Các dịch vụ của nội thất Đăng Khôi bao gồm: sửa đồ gỗ, làm gác xép, sửa sàn gỗ...
ReplyDeleteThnkyou for nice tutorial blockchain training
ReplyDeletemicrosoft azure certification career has a good hope
ReplyDeleteI thank you for your post hadoop certification
ReplyDeleteI was searching for exactly the same information.Thanks for sharing.Good work.Keep it up.These days Big data is trending technology.“Without big data, companies are blind and deaf, wandering out onto the web like deer on a freeway.”If you are looking for any online courses on big data visit our site.
ReplyDeleteBig Data Hadoop Online Training Courses
This post is really nice and informative. The explanation given is really azure tutorial comprehensive and informative...
ReplyDelete
ReplyDeleteThis is most informative and also this post most user friendly and super navigation to all posts. Thank you so much for giving this information to me. Hadoop training in Chennai.
Java training in chennai | Java training in annanagar | Java training in omr | Java training in porur | Java training in tambaram | Java training in velachery
The only blog I have is on Scrapbook.com, is there anyway to enter without one.thanks a lot guys.
ReplyDeleteC and C++ Training Institute in chennai | C and C++ Training Institute in anna nagar | C and C++ Training Institute in omr | C and C++ Training Institute in porur | C and C++ Training Institute in tambaram | C and C++ Training Institute in velachery
Really nice and interesting post. I was looking for this kind of information and enjoyed reading this one. Keep posting. Thanks for sharing.
ReplyDeletedata science course in guntur
Nice and good article. It is very useful for me to learn and understand easily. Thanks for sharing your valuable information and time. Please keep..
ReplyDeleteMicrosoft Windows Azure Training | Online Course | Certification in chennai | Microsoft Windows Azure Training | Online Course | Certification in bangalore | Microsoft Windows Azure Training | Online Course | Certification in hyderabad | Microsoft Windows Azure Training | Online Course | Certification in pune
Good Post! , it was so good to read and useful to improve my knowledge as an updated one, keep blogging.After seeing your article I want to say that also a well-written article with some very good information which is very useful for the readers....thanks for sharing it and do share more posts likethis
ReplyDeletehttps://www.3ritechnologies.com/course/online-python-certification-course/
Good blog post,
ReplyDeleteDigital Marketing Course in Hyderabadls
ReplyDeleteThat is nice article from you , this is informative stuff . Hope more articles from you . I also want to share some information about Pet Dentistry in vizag
binance güvenilir mi
ReplyDeleteinstagram takipçi satın al
takipçi satın al
instagram takipçi satın al
shiba coin hangi borsada
shiba coin hangi borsada
tiktok jeton hilesi
is binance safe
is binance safe
that is really an great web Mobile Prices Bangladesh
ReplyDeleteHelpful data are extremely intriguing these days on the web. I might want to thank the writer who composed and delivered this article. The Turkish government provides electronic Turkey visa USA which is travel authorizations to United States citizens. It is for people who want to travel to Turkey without applying for a visa in advance and is different from a standard visa.
ReplyDeleteWhile I have read your article several times, I find many valid points in it. I am confident your readers will enjoy it. Travelers can apply for an e visa Turkey which is very easy. If you have an internet connection and valid documents, you can apply online from anywhere in the world.
ReplyDeleteVery nice post. I just came across your blog and wanted to say that I love browsing your blog posts. e visa online, E Visa application process now is online. You can e visa apply online within 5 to 10 minutes you can fill your visa application form. And your visa processing depends on your nationality and your visa type.
ReplyDelete
ReplyDeleteLovely post..! I got well knowledge of this blog. Thank you!
Solicitation Of A Minor VA
Online Solicitation Of A Minor
VNC Connect Enterprise 6.11 Crack is easy-to-use remote access software that lets you connect to a remote computer anywhere in the world,.Vnc Viewer Plus License Key
ReplyDeleteis it ok if buy a macine for somewhere on website saying bending machine for sale ??
ReplyDeleteWe greatly value the information you gave us. Thank you for sharing with us. The Ten Most LGBTQ Friendly Countries represent beacons of progress and inclusivity in today's global landscape. These nations have made significant strides in promoting and protecting the rights of the LGBTQ community.
ReplyDeleteWonderful , really it is a very helpful post. I heartily appreciate your efforts and I will be waiting for your further posts. Thank you once again. If you're a Jamaican citizen aiming to visit India, here's what you need. India Visa for Jamaica Citizens. Go online, complete the form, attach required documents, and pay the fee. Wait a bit for processing. Once approved, you're ready to explore India. Adequate preparation ensures a seamless online visa process.
ReplyDeleteSafePal is a secure decentralized wallet that enables users to import, recover and manage wallets and crypto-assets on mobile devices. To create a SafePal software wallet, you will need to download the SafePal app, set up a security password, and create a new wallet. The wallet will be generated a 12 or 24-word mnemonic phrase, which is the master key to your wallet. It is important to keep this phrase safe and secure.
ReplyDeleteHii Everyone, Welcome to the hassle-free way of traveling to Turkey! Say goodbye to lengthy procedures and queues. Apply for your Turkish e-Visa online today and experience the convenience of obtaining your travel authorization from the comfort of your home.
ReplyDeleteHello! I'd like to extend my heartfelt appreciation for the invaluable information shared on your blog. vfs global Saudi arabia Your gateway to seamless visa and passport services. Explore efficient solutions for travel, study, work, and more with our expert assistance.
ReplyDeleteIn a sea of digital noise, your work emerges as a guiding star of relevance and substance. India Opens Doors to Saudi Arabian Citizens, extending an invitation to explore its vibrant culture, diverse landscapes, and historical treasures. This gesture enhances cultural exchange and strengthens ties between the two nations, fostering mutual understanding and cooperation. Saudi Arabian travelers can now embark on enriching journeys across India, discovering its beauty and hospitality.
ReplyDeleteHello everyone, Navigating the requirements for an Zurich transit visa for Indian passport holders is crucial for a smooth journey. Understanding this process ensures a hassle-free stopover in India for travelers with Zurich passports.
ReplyDeleteYour unwavering commitment to your blog is truly admirable. It's clear that you possess a genuine ardor for your chosen topics, and your writing seamlessly marries expertise with entertainment. Keep up the exceptional work, and I'm eagerly anticipating delving into your forthcoming posts. Here's to your ongoing triumphs! Cheers!
ReplyDeleteHow To Fix Trust Wallet Balance Not Showing
A Step-by-Step Guide to Resetting Your Password on the Crypto.com Exchange
Hello! Your support is invaluable, motivating us to provide even more engaging content. Thank you for being our inspiration and encouraging us to continue creating. To track your Indian Passport Tracking in Saudi Arabia visit the official Indian government's passport tracking website. Enter your file number and date of birth to get real-time updates on your passport status.
ReplyDeleteYour blog has a unique way of sparking introspection. Your recent post resonated deeply, and I've been discussing its ideas with friends ever since. Your thought-provoking content is a gem. India and Uganda share strong and growing ties between India and Uganda, symbolizing a promising future of collaboration and mutual growth between the two nations.
ReplyDeleteThis comment has been removed by the author.
ReplyDeleteHello, kindred spirit! Your blog consistently brightens my day. Your engaging style and informative content are a perfect blend. This post was no different; it felt like a warm chat with a friend. Thanks for making knowledge feel approachable and friendly. Cameroon Cameroon Visa refusal by Philippines: Travelers may encounter obstacles when seeking entry, underscoring the need for diligent research and preparation for smoother visits.
ReplyDeleteI appreciate the variety of topics you cover on your blog. Your recent post was insightful, and I look forward to more engaging reads. Explore the Turkey Visa for Bhutan: application steps, requirements, and essential details for travelers.
ReplyDeleteInteresting topics covered here. Well-written and informative content. Keep up the good work and bring more engaging content! Visa thrives in post-pandemic travel rebound period as global mobility increases, highlighting the importance of secure and efficient payment solutions.
ReplyDelete
ReplyDeleteYour post is a gem in my daily blog explorations. While I typically don't read articles on blogs, your write-up has inspired me to make an exception. Your writing style is truly captivating, and I have to say, this article is exceptionally well-crafted. Thank you for presenting this outstanding piece.how much is Turkey visa fee 2024? The Turkey visa fee for 2024 may vary depending on your nationality, type of visa, and duration of stay.
Within your blog post, an intellectual masterpiece comes to life—an eloquent symphony of profound insights. Each paragraph is a brushstroke, crafting a vivid tapestry within the intricate landscapes of knowledge. Your adept navigation through complexity is commendable, creating a harmonious blend of intellect and expression. i want to share some information with you You can conveniently perform a Turkey visa check online by passport number. This streamlined process allows you to verify your visa status effortlessly, ensuring a smooth travel experience for your journey to Turkey.
ReplyDeleteThis ultimate guide of how to buy Pepe Meme Coin is a game-changer for anyone looking to explore the world of meme-inspired cryptocurrencies. The detailed step-by-step instructions provided here make the buying process a breeze, ensuring even newcomers can navigate the crypto space with confidence. Pepe Meme Coin brings a refreshing and humorous perspective to the market, making it an exciting addition to any crypto portfolio.
ReplyDeleteWithdraw Money from BlockFi Wallet is a straightforward process that ensures you have access to your funds whenever you need them. Start by logging into your BlockFi account and navigating to the withdrawal section. From there, select the cryptocurrency you want to withdraw and enter the amount. Make sure to double-check the withdrawal address to ensure your funds are sent to the right destination. Once you've confirmed all the details, submit your withdrawal request and wait for it to be processed. BlockFi provides a user-friendly platform that makes it easy to manage your crypto assets and access your funds whenever you need them.
ReplyDeleteIf you need to contact Curve Exchange Support for assistance or inquiries, there are several ways to reach out for help. You can visit Curve's official website and explore their support section, where you'll find helpful resources such as FAQs and guides. Additionally, Curve maintains an active presence on social media platforms like Twitter and Discord, providing opportunities for users to engage with the community and seek assistance from Curve representatives. Whether you have questions about using the platform or encounter any issues, reaching out to Curve Exchange Support through these channels can help you get the support you need promptly.
ReplyDeleteTo know How do I Contact PancakeSwap Support, you can visit their official website and look for the "Support" or "Contact Us" section. There, you should find information on how to reach their customer service team. Additionally, you may be able to contact them through their official social media accounts by sending direct messages or leaving comments. Make sure to use verified channels to avoid potential scams or phishing attempts.
ReplyDeleteIt can be frustrating when your Coinbase Wallet Not Showing Balance accurately. This issue can arise due to several reasons, such as network congestion, pending transactions, or account verification processes. However, there's no need to panic. By staying calm and exploring potential solutions, like waiting for transactions to confirm or ensuring your account is fully verified, you can often resolve the problem. If you're still experiencing difficulties, don't hesitate to reach out to Coinbase Support for assistance. They can provide personalized help to get your balance back on track.
ReplyDeleteIf you need to Contact MetaMask customer support, the best way is to visit their official website and head to the support section. From there, you can find helpful resources such as FAQs and troubleshooting guides. If your issue isn't resolved through these resources, you can usually find a way to contact their support team directly. Make sure to provide clear and detailed information about your problem to expedite the resolution process.
ReplyDeleteWhen it comes toSpeed Up or Cancel Pending Ethereum Transactions on Ledger Wallet, there are a few avenues users can explore. While Ledger Live doesn't offer direct support for features like "Replace by Fee" (RBF), individuals can turn to alternative wallets like MyEtherWallet (MEW) or MyCrypto. By importing their Ledger wallet using the recovery phrase, users gain access to the pending transaction. They can then adjust the gas fee to accelerate its processing or create a new transaction with a higher fee to supersede the original one. However, users should exercise caution and verify all transaction details and fees meticulously to mitigate any potential risks associated with cryptocurrency transactions.
ReplyDeleteTo Contact Coinbase Web3 Wallet, you can follow these simple steps. First, ensure you have both your Coinbase account and Web3 Wallet open. Then, navigate to the settings or wallet management section of your Coinbase account. Look for an option related to connecting a Web3 wallet or adding a new wallet. Click on it, and you'll likely be prompted to scan a QR code or input a specific address. In your Web3 Wallet, find the option to connect to an external service or add a new wallet.
ReplyDeleteTake a simple guide to transfer funds from Argent to MetaMask. Open your Argent app and access the Transfer menu within your zkSync Era account. Next, open your MetaMask wallet, click on the circle in the top-right corner to access the settings, and copy your MetaMask public address. Return to the Argent app, enter the copied MetaMask address as the recipient, and confirm the transfer to securely move your assets from Argent to MetaMask.
ReplyDeleteTo resolve the "Swap Service is Currently Unavailable" issue in Atomic Wallet, ensure your app is updated to the latest version, as updates often fix bugs and improve functionality. Check your internet connection to ensure it's stable, as network issues can affect the swap service. If the problem persists, try clearing the cache and restarting the app. Additionally, visiting Atomic Wallet’s official support page or contacting their
ReplyDeleteTo resolve a 0 Coinbase Showing 0 Balance, first ensure that your app is updated to the latest version, as updates can fix display bugs and improve functionality. Clear your app's cache and restart the application to refresh your account data. Check your internet connection to ensure it's stable and that the Coinbase servers are not experiencing downtime. If the issue persists, log in to your account on the Coinbase website to verify your balance; this can help determine if it's an app-specific problem. For further assistance, contact Coinbase support to report the issue and receive guidance on troubleshooting steps.
ReplyDeleteGreat Information! Checked it out and left my thoughts. If you're interested in Professional Crypto Customer Care Advisor feel free to explore How Can I Put PayPal money into my Apple Pay account? insights.
ReplyDeleteGreat Information! Checked it out and left my thoughts. If you're interested in Professional Crypto Customer Care Advisor feel free to explore How do I add my bank card to Apple Pay insights.
ReplyDeleteTo Transfer Money From Crypto.com To Bank Account, first ensure that you have completed the necessary KYC (Know Your Customer) verification on the Crypto.com platform. Next, link your bank account to your Crypto.com account by navigating to the "Accounts" section, selecting "Fiat Wallet," and then choosing "Withdraw." Enter the amount you wish to transfer and select your linked bank account. Confirm the transaction details and initiate the transfer. The funds should be deposited into your bank account within a few business days, depending on your bank's processing times.
ReplyDeleteIf your coinbase account restricted, it could be due to various reasons, such as unusual account activity, failure to complete verification, or violating Coinbase's terms of service. To resolve this issue, log into your account and check for any notifications or requests for additional information. Completing any pending verifications or updating your information might help restore access. If the issue persists, contact Coinbase support for assistance. They can guide you through the necessary steps to lift the restriction and regain full access to your account.
ReplyDeleteA Transaction in Coinbase Wallet involves sending or receiving cryptocurrency directly through the wallet interface. Users can seamlessly manage their assets, execute trades, or interact with decentralized applications (dApps). The wallet provides a secure and private environment for transactions, ensuring that users maintain control over their private keys and funds. With Coinbase Wallet, users can easily access their digital assets and perform transactions without needing to rely on third-party services, making it a versatile tool for managing and using cryptocurrencies.
ReplyDeleteYou'll first need to link a common bank account to both apps to transfer money from Venmo to Cash App. Transfer the desired amount from your Venmo balance to your linked bank account. Once the funds appear in your bank account, you can then transfer them from the bank to your Cash App balance. This process typically takes one to three business days for each transfer. While there isn't a direct way to transfer money between Venmo and Cash App, a linked bank account provides an efficient workaround.
ReplyDeleteTo recover your MyEtherWallet (MEW) account, you must use your private key, recovery phrase, or Keystore file. If you have these, visit the MEW website, select "Access My Wallet," and follow the prompts. These recovery details are necessary for access to be recovered, as MEW cannot restore accounts for security reasons. Always store your recovery information securely.
ReplyDeleteTo Transfer Money From Crypto.com To Bank Account, first, sell your crypto for fiat in the app. Then, go to the "Transfer" option, select "Withdraw," choose "Fiat," and pick your bank account. Enter the amount to transfer and confirm. The funds should arrive in your bank within 3-5 business days.
ReplyDeleteIf your coinbase balance not showing, first check if the app or website is up to date. Ensure you're connected to the internet and try refreshing the page or restarting the app. If the problem persists, clear your browser cache or app data. You can also check Coinbase's status page for outages or contact their support for further assistance.
ReplyDeleteTo Withdraw Money from Crypto DeFi Wallet, connect the wallet to a compatible decentralized exchange (DEX). Select the cryptocurrency you wish to convert to fiat, execute the swap, and transfer the converted funds to a centralized exchange. From there, you can withdraw the money to your bank account. Always double-check wallet addresses to avoid errors.
ReplyDeleteIf your Coinbase Account Under Review, it may be due to suspicious activity, verification issues, or security concerns. During this period, account access might be limited. To resolve the issue, check your email for instructions from Coinbase, and provide any requested information. Contact Coinbase support for further assistance, and ensure your account details are up to date.
ReplyDeletehttps://saadwmsblog.blogspot.com/2019/01/blockchain-in-supply-chain-reality-or.html?sc=1724843211792#c4562775042281553878
ReplyDeleteIf your coinbase balance not showing, it could be due to a temporary glitch, account synchronization issues, or an update delay. Ensure you're using the latest app version and try refreshing the page or logging out and back in. If the problem persists, contact Coinbase support for further assistance to resolve the issue and access your funds.
ReplyDeleteIf your Coinomi wallet is experiencing a "coinomi no connection" issue, ensure your internet connection is stable and try restarting the app. Clear the app's cache, update to the latest version, or check if the Coinomi server is down. If the problem persists, consider switching networks or reinstalling the app for a fresh start.
ReplyDeleteTo Transfer Money From Crypto.com To Bank Account, first, sell your cryptocurrency for fiat currency within the app. Once the funds are in your fiat wallet, select the "Withdraw" option and choose your linked bank account. Enter the amount you'd like to transfer, confirm the details, and submit the request. The transfer process typically takes 2-5 business days, depending on your bank’s processing times.
ReplyDeleteIf your Coinbase Account Under Review, it typically means the platform is conducting a routine check to ensure compliance with its security policies. Unusual activity or verification requirements can trigger reviews. During this period, certain account functions may be temporarily restricted. Contact Coinbase support for more information or wait for the review process to be completed.
ReplyDeleteIf your coinbase balance not showing, it could be due to a temporary glitch or an issue with account synchronization. Ensure that your app or browser is updated, and try refreshing your account. If the problem persists, contact Coinbase support to investigate the issue and restore your balance visibility quickly.
ReplyDeleteIf your Base ETH Not Showing in MEW Wallet, first double-check that you're connected to the correct network (Mainnet) and logged into the right account. Sometimes, switching networks and back again helps refresh the wallet balance. You can also try clearing your browser cache or using a different browser. If that doesn't work, check your balance on Etherscan to ensure the ETH is still associated with your address. If it's visible there but not in MEW, the issue is likely display-related in the wallet.
ReplyDeleteTo Fix Internal JSON-RPC Error in Metamask first try clearing your browser's cache or the app’s cache if you're using the mobile version. Ensure MetaMask and your browser are updated to the latest versions. If the issue persists, switch networks or reset your MetaMask account under settings. This won’t affect your funds but may resolve the error.
ReplyDeleteTo Withdraw Money from Crypto DeFi Wallet, first, connect the wallet to a centralized exchange that supports your cryptocurrency. Transfer the desired amount from your DeFi wallet to the exchange, where you can sell your crypto for fiat currency. Once sold, initiate a withdrawal to your linked bank account. Always double-check transaction details to avoid errors.
ReplyDeleteWithdraw from Rainbow Wallet is simple and secure. Open the app, select the cryptocurrency you want to withdraw from the "Assets" tab, and tap "Send." Input the recipient's wallet address or scan their QR code, then specify the amount to transfer. Review all transaction details, including any associated fees, and confirm. Once confirmed, the transaction will be processed on the blockchain, and the recipient will receive the funds after validation.
ReplyDeleteA Coinbase Account Under Review typically means that the platform is conducting a routine check to ensure compliance with regulatory standards or to verify account information. This process may temporarily restrict access to certain features, such as withdrawals or trading. Reviews can be triggered by unusual activity, large transactions, or verification updates. It's advisable to monitor your email for updates from Coinbase during this period.
ReplyDeleteContact Blockchain Customer Service by calling our team for wallet assistance, transaction troubleshooting, deposit and withdrawal guidance.
ReplyDeleteIf Coinomi shows "coinomi no connection available, " check your internet connection and ensure it’s stable. Try switching between Wi-Fi and mobile data or restarting your device. If the issue persists, clear the app’s cache or reinstall it. Make sure Coinomi is updated to the latest version. Lastly, check if the Coinomi servers are experiencing downtime or maintenance.
ReplyDeleteHow to Transfer from MetaMask to KuCoin, start by logging into your MetaMask wallet and selecting the correct network (e.g., Ethereum). Then, log into KuCoin and go to the "Deposit" section to find your KuCoin wallet address for the specific cryptocurrency. Copy that address, return to MetaMask, and click "Send." Paste the KuCoin address in the recipient field, enter the amount, and confirm the transaction. Double-check all details, including the network, before submitting. The transfer will be complete after the transaction is confirmed on the blockchain.
ReplyDeleteIf your deposit Stuck on Pending in Coinbase Wallet, it's likely due to network congestion or low gas fees. To speed it up, you can try increasing the gas fees if the option is available. Otherwise, you might need to wait for the network to process the transaction. You could also attempt to cancel the pending deposit by sending a new transaction with the same nonce and higher gas fees. Make sure you have enough funds to cover the necessary gas fees, as this is often the key reason for delays.
ReplyDeleteTo Transfer Money from Venmo to Cash App, you’ll need to link a mutual bank account to both apps. First, transfer your funds from Venmo to your linked bank account by selecting “Transfer to Bank” in Venmo and choosing the bank account. Once the funds are available in your bank account, open Cash App, tap “Add Cash,” and transfer the desired amount from your bank to Cash App. Unfortunately, there’s no direct transfer option between Venmo and Cash App, so using a bank account is necessary.
ReplyDeleteHow To Send USDT From Trust Wallet To Binance is easy. First, make sure you have USDT in your Trust Wallet. On Binance, go to "Wallets," select USDT, and click "Deposit." Copy the provided deposit address, ensuring you choose the correct network like BEP20 or ERC20. In Trust Wallet, click "Send," paste the Binance deposit address, and specify the amount of USDT. Confirm the details and complete the transfer. Your USDT should arrive in your Binance account after processing.
ReplyDeleteLearn how to Withdraw Tether from Coinbase is straightforward. First, log in to your Coinbase account and go to the “Assets” section. Select USDT from your list of holdings and click “Send.” Enter the recipient wallet address, ensuring it matches the blockchain type (e.g., ERC-20 or TRC-20). Input the amount to send, review the transaction details, and confirm.
ReplyDeleteThe Coinbase Web3 Wallet is a powerful self-custodial wallet designed to securely and seamlessly manage digital assets. It enables users to interact with DeFi platforms, NFTs, and decentralized applications (dApps) while maintaining full control over their private keys. With multi-chain support and robust security features, the wallet ensures a safe and user-friendly experience. Available as a mobile app and browser extension, it’s an ideal choice for exploring the Web3 ecosystem.
ReplyDeleteBitstop ATM provides customer support to assist users with transactions, troubleshooting, and other inquiries. While they offer Bitstop ATM Customer Service through various channels, including email and phone support, availability of live chat support may vary. If you're looking for immediate assistance, visiting their official website or contacting their support team directly is the best way to confirm live chat options. Users experiencing issues with Bitcoin purchases, withdrawals, or technical problems can reach out for prompt assistance.
ReplyDeleteLearn how to transfer funds from Coinbase to Kraken quickly and securely. Follow our step-by-step guide to move your crypto assets, avoid fees, and ensure a smooth transaction between exchanges. Start your transfer today!
ReplyDeleteGate io Customer Care support to assist users with inquiries related to trading, account security, deposits, withdrawals, and other platform services. Users can reach Gate.io’s customer care through their official website by submitting a support ticket, using the live chat feature, or accessing the help center for FAQs and troubleshooting guides.
ReplyDeleteLearn how to buy Solana with a credit card safely and instantly using Kraken. This step-by-step guide covers why Solana (SOL) is a great investment, the best ways to purchase it, and secure storage options. Start your crypto journey today with fast transactions, low fees, and top-tier security. Buy SOL now!
ReplyDeleteCrypto.com KYC verification can fail for a few reasons, and it's honestly frustrating when it happens. Common issues include blurry or low-quality photos of your ID, submitting expired documents, mismatched information between your ID and your account details, or even minor mistakes like typos. Sometimes, system errors or high traffic on their platform can also cause delays or unexpected rejections.
ReplyDeleteRead More: Why does Crypto.com KYC Verification keep failing
Facing a "Crypto.com withdrawal failed" problems? Discover expert tips, common causes, and solutions from KrakenWallets.info to resolve the issue quickly and securely. Crypto Investment Customer Service Phone Number.
ReplyDeleteIf you're trying to know how to Transfer from Keystone to another account, the process is pretty straightforward. Use the Keystone companion app to set up the transaction by entering the recipient’s address and amount. Then scan the QR code with your Keystone device to review and sign it securely. Once signed, scan it back into the app to broadcast the transaction.
ReplyDelete